Invasive mammals such as rats are associated with negative impacts on native fauna and their eradication on islands around the world has been proved to result in substantial conservation gains. Fernando de Noronha is an archipelago located off the north-east coast of Brazil and has records of native fauna negatively impacted by introduced rats. We used D-Loop sequences and 14 microsatellite markers to investigate gene flow between three populations of Rattus rattus across two islands of Fernando de Noronha. Using both methods we found very great differentiation between the two islands, indicating very low or no gene flow. Our findings suggested if Rata Island were cleared of rats it would have a low probability of reinvasion occurring from the main island. Thus, we recommend an eradication program be implemented on Rata Island following the successful program on nearby Meio Island.
Marine islands represent 5.3% of Earth’s land and hold 19% of bird species, 17% of rodents and 17% of flowering plants of the world, which means islands are proportionally richer in species than continental areas. However, this high diversity in insular areas has suffered 30 times more extinctions and has 14 times more critically endangered species per km2 than continental areas (Tershy et al., 2015). Invasive mammals are considered as one of the major factors responsible for biodiversity loss on oceanic islands (Caujapé-Castells et al., 2010), and rats are especially significant given their commensal behaviour with humans that allows them to travel long distances and their omnivorous diet that makes them able to exploit many habitats (Drake and Hunt, 2009; Varnham, 2010).
In 2015, more than 580 islands had successfully executed rat eradication worldwide (Russell and Holmes, 2015) and many native species, including invertebrates, birds, mammals and reptiles, are shown to be benefited after invasive mammal eradication (Jones et al., 2016). On Fernando de Noronha there are records of rats preying on turtle nests, and the most common species of invasive rat on the island, the black rat (Rattus rattus) (Russell et al., 2018), was possibly one of the causes for the extinction of Noronhomys vespuccii, an endemic rodent from Fernando de Noronha (Carleton and Olson, 1999). Besides that, Noronha holds 11 of 14 seabird species occurring on Brazilian offshore islands, some almost restricted to rat-free secondary islets and with small populations that could become extinct soon (Mancini et al., 2016; Russell et al., 2018) and therefore are vulnerable in case of rat invasion.
Rat eradication on Fernando de Noronha would be a complex task but also has the potential to bring significant benefits to local biodiversity and human health (Russell et al., 2018). Russell et al. (2018) recommended that Rata Island could be used as an experimental site for pest eradication to protect and restore breeding seabird colonies. To minimize eradication chances of failure, an increasing number of studies have recommended to perform pre-eradication genetic analysis to define appropriate eradication units (e.g. Abdelkrim et al., 2005; Savidge et al., 2012; Adams et al., 2014). As an example, a genetic assessment of the R. rattus population on Pearl Island, New Zealand, revealed that an eradication failure was most likely due to reinvaders from an adjacent population (Russell et al., 2010). If the genetic assessment had been performed before the eradication operation this migration between islands might have been more obvious. On the other hand, Adams et al. (2014), performed a pre-eradication genetic analysis on the common brushtail possum (Trichosurus vulpecula) population in Dunedin and on the Otago Peninsula, New Zealand, and detected a potential reinvasion pathway. In that case, the authors recommended that the Eastern Peninsula be treated as one eradication unit. The definition of eradication units can help to decide how biosecurity effort may be applied to prevent recolonization from surrounding islands after an eradication is done as one unit.
On Fernando de Noronha the two largest islands (the main island and Rata Island) are separated by 1800 m which is more than black rats are believed to be able to swim (Russell and Clout, 2005) and hence recolonize. However, between those islands there are three smaller islands (Rasa, Sela Ginete and Meio) that might be used as stepping stones and enable gene flow across the islands. This paper aims to investigate gene flow between two putative populations on the main island and a third on Rata Island, assessing population genetics of the most common invasive rat (R. rattus), and validating the feasibility of the Rata Island eradication strategy proposed by Russell et al. (2018).
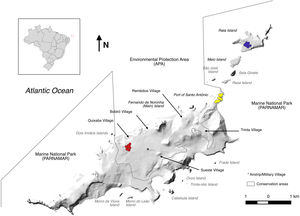
Material and methodsStudy areaFernando de Noronha is a volcanic archipelago lying 345 km off the north-east coast of Brazil (3°50′S, 32°26′W), composed by 21 islands with total land area of 26 km2 (Fig. 1). The main island is 17 km2, and is the only inhabited. The island has a human population around 3,000 residents and receives more than 90,000 tourists every year (MTUR, 2017). About 70% of the main island is a National Park, while the rest is classified by law as environmental protection area, where infrastructure is permitted but controlled. The remaining islands can only be visited under permits of the Institute Chico Mendes for Biodiversity Conservation (Silva, 2013a).
Our area of interest was the north-east end of the archipelago comprising the main island and stepping stones of Rasa, Sela Ginete, Meio and Rata Islands (Fig. 1). Rasa Island (5 ha) is low and flat and almost connected to the main island at low tide, but with undercut rock and strong currents remains difficult to access. Sela Ginete (5 ha) is a tall sparsely vegetated granitic column and also treacherous to land upon. Meio Island (17 ha) is low, flat and undercut also like Rasa Island, and in 2017 black rats were successfully eradicated (unpublished data) as a proof of concept for the application of rat eradication methodologies on Fernando de Noronha. Rata Island (89 ha) is a combination of undercut rock but with a main rocky beach exposed to a strong swell, and the forested island gently rises up to a high point (Fig. 2).
Sample collectionBlack rat tissue samples were collected from three sites across the archipelago from 2015 to 2018 which we a priori treated as population units. Rats were captured in live traps (Tomahawk, 21 × 21 × 30 cm) and euthanised for tissue collection using Ketamine-Xylazine (K: 90 mg/kg + X: 10 mg/kg SQ) for general anaesthesia, followed by euthanasia using 0.2 ml of T-61 (Embutramide 200 mg/ml + Mebezonium iodide 50 mg/ml + Tetracaine hydrochloride 5 mg/ml). We were most interested in connectivity from the north-east point of the main island, around the port where rats might either depart by swimming along the stepping stones of the north-east chain, or by hitch hiking on boats. Therefore, the first sampling site was the harbour on the main island. For comparison of genetic diversity across the main island rats were also sampled from the village of Quixaba lying in the middle of the main island 4.8 km from the port. At last, rats were sampled on Rata Island. We were not able to obtain samples from Sela Ginete due to its inaccessibility, and no samples were available from Meio Island prior to the rats being eradicated.
DNA extraction, genotyping and sequencingGenomic DNA was extracted from tails and ears tissue samples, preserved in ethanol using DNeasy Blood and Tissue Kit (Qiagen), following the manufacturer’s protocol.
D-Loop mitochondrial region was amplified using EGL4L and RJ3R primers developed by Robins et al. (2007). Each PCR reaction had a final volume of 20 μL, 10 mM Tris HCl pH 8.3; 50 mM KCl, 2.5 mM MgCl2, 0.5 μM of each primer, 0.15 mM dNTPs, 0.5 U of Platinum Taq DNA polymerase (Invitrogen) and 1 μL of DNA template. Thermocycler (Eppendorf Mastercycler pro S) conditions were: initial denaturation step of 94 °C for 2 min; 35 cycles of 94 °C for 30 s, 60 °C for 30 s and 72 °C for 1 min and final extension step of 72 °C for 5 min. PCR products were purified with ExoSAP-IT (Affymetrix, Inc.). Sequencing was carried out at the University of Auckland DNA Sequencing Facility, New Zealand, using the BigDye Terminator version 3 sequencing kit and a capillary ABI3130XL (Applied Biosystems) DNA automated sequencer.
We used 14 microsatellite markers to genotype the populations. Eleven markers were developed by Jacob et al. (1995) from Rattus norvergicus, but have been used for R. rattus by many authors (e.g. Abdelkrim et al., 2010; Brouat et al., 2014; Pichlmueller and Russell, 2018): D2Rat234, D11Mgh5, D5Rat83, D7Rat13, D10Rat2, D12Rat76, D15Rat77, D16Rat81, D18Rat96, D19Mit2 and D20Rat46. Three other markers specifically designed for R. rattus characterized by Loiseau et al. (2008) (Rr14, Rr17, Rr114) were used. Markers were chosen from different chromosomes to avoid physical linkage. Forward primers were labelled with fluorescent dyes: 6-FAM, HEX, VIC and PET (Applied Biosystems) (Supplementary Material 1).
The polymerase chain reactions (PCR) were performed as multiplex reactions in 10 μl volumes containing 1 μl of extracted DNA (>40 ng DNA/ μl), 1 X QIAGEN Multiplex PCR Master Mix and 0.2 μM of each primer used in the reaction. Loci were multiplexed in sets of two to seven. The thermocycler conditions were: initial heat activation of 95 °C for 15 min, 30 cycles of 94 °C for 30 s, 55 °C for 90 s and 72 °C for 60 s, and final extension of 72 °C for 20 min.
PCR products were diluted 10-fold, then 1 μL of the PCR solution was mixed with 0.4 μL of GeneScan™ 600 LIZ® (Thermo Fisher Scientific) and 10 μL of Hi-Di Formamide (Applied Biosystems), this mixture was submitted to heatshock treatment, 95 °C for 5 min, 4 °C for 5 min. Genotyping runs were performed on an ABI3130XL (Applied Biosystems) automated sequencer and analysed using GENEIOUS Prime 2019.0.4 (https://www.geneious.com) including the Microsatellite Analysis External Plugin version 1.4.6 (Biomatters Ltd.). To ensure scoring accuracy we re-genotyped 5% of samples.
Data analysisFor D-Loop, all the raw sequences were aligned in GENEIOUS Prime 2019.0.4 adjusted manually and trimmed to a common length of 520 bp.
The software Micro-Checker version 2.2.3 (Van Oosterhout et al., 2004) was used to test for the presence of null alleles, using Bonferroni correction and 3000 randomisations.
The number of alleles (NA), allelic richness (Ar), observed heterozygosity (HO), and within population gene diversity (HS) were calculated with FSTAT 2.9.4 (Goudet, 1995) for each population and all individual loci (Table 2). FIS and FST values were calculated according to Weir and Cockerham (1984). Pairwise FST values between populations (Table 3) were calculated and p-values were obtained with 95% confidence intervals, after 3,000 permutations. A test for departure from Hardy-Weinberg equilibrium was also carried out with FSTAT using 21,000 randomisations.
To approach the question of spatial genetic structure we use Bayesian clustering in software STRUCTURE 2.3.4 (Falush et al., 2003; Pritchard et al., 2000). The admixture model with correlated allele frequencies was run with 200,000 Markov chain Monte Carlo iterations for 10 runs, after a burn-in period of 1,000,000 and number of clusters ranging K = 1–4. The output file from STRUCTURE was used in STRUCTURE HARVESTER v0.6.94 (Earl and vonHoldt, 2012) to calculate ΔK values (the rate of change in the log probability of data between successive K values) as suggested by Evanno et al. (2005). Individual membership assignments estimated in STRUCTURE were aligned by CLUMPP 1.1.2 (Jakobsson and Rosenberg, 2007) and DISTRUCT 1.1 (Rosenberg, 2004) was used to generate the bar plot.
We also perform principal components analysis (PCA) and use the saddlepoint approximation method described by McMillan and Fewster (2017) to visualize the genetic assignment of each sample. This analysis was performed in R 3.5.2 (R Core Team, 2017) with the package geneplot v0.1.0 (McMillan and Fewster, 2017).
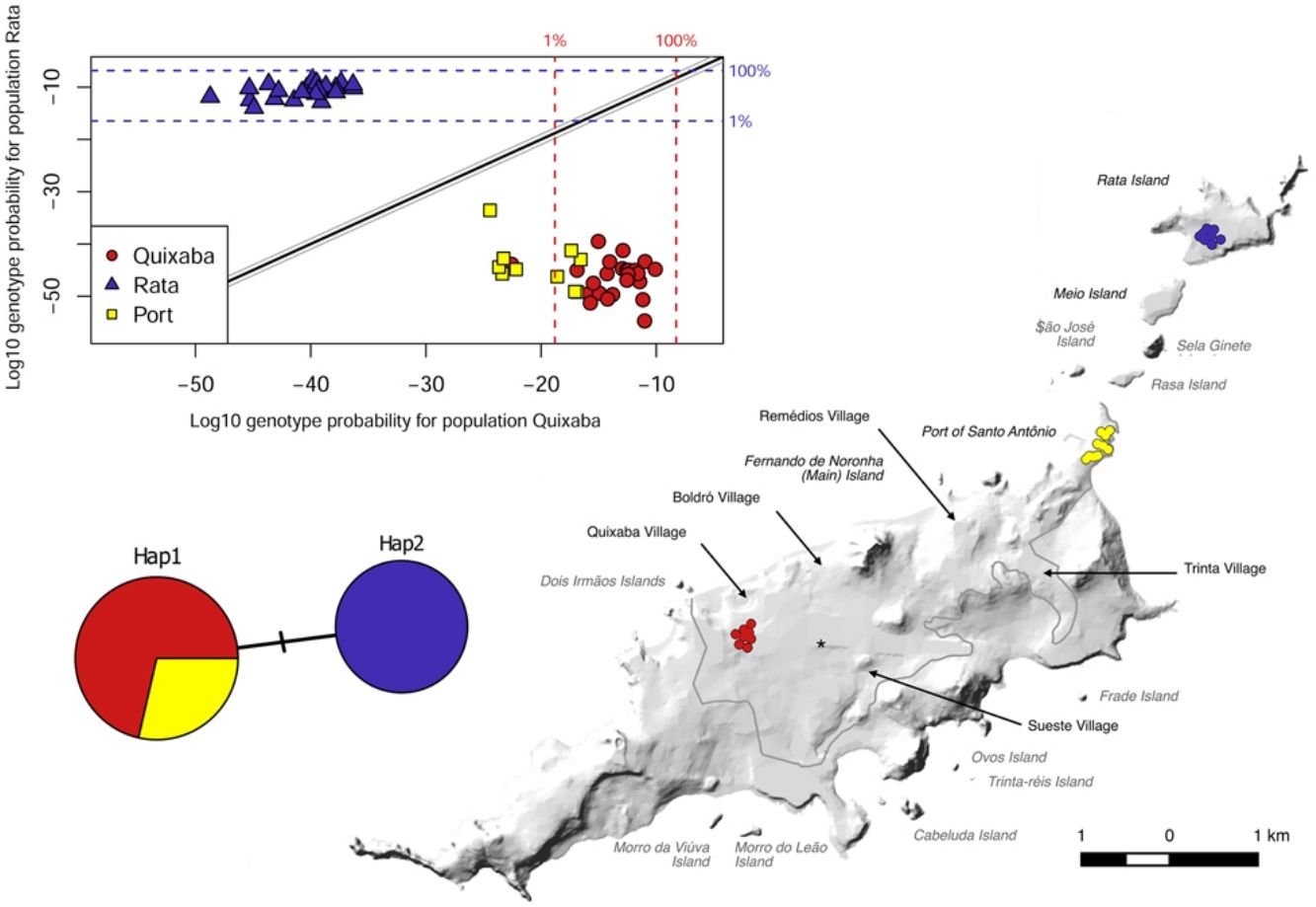
ResultsD-LoopA total of 58 individuals were caught and included in our study: 35 from the main island (25 from Quixaba village and 10 from the Port area) and 23 from Rata Island. The sequencing of D-Loop region for the 58 individuals resulted in only 2 haplotypes that were geographically partitioned. Hap1 was found in all 35 individuals from the main island (Quixaba and Port populations) and Hap2 was found only in individuals from Rata. The haplotypes differs from each other by only one base pair transition C↔T. Using the mitochondrial reference genome for R. rattus described by Robins et al. (2008) (GenBank NC_012374) this transition is located at base position 15,757. The two haplotypes were deposited in GenBank (Accession numbers MN746385 and MN746386).
MicrosatelliteThe genotyping success was 99.7%. As expected, the population from Rata had lower diversity than the Quixaba population regarding number of alleles, allelic richness, gene diversity and observed heterozygosity (Table 1). Even the Port population, that had only 10 individuals genotyped, had higher allelic richness than the Rata population and the number of alleles was only slightly higher for Rata.
Sampling location, number of sampled individuals (NS), number of alleles (NA), allelic richness (Ar), private alleles (PA), within population gene diversity (HS), and observed heterozygosity (Ho; derived from HS and FIS values as Ho = HS−(FIS × HS)), FIS values (not significant, based on 42,000 randomisations).
| Population | Ns | Na | Ar | PA | Hs | Ho | FIS |
|---|---|---|---|---|---|---|---|
| Quixaba | 25 | 86 | 4.7 | 27 | 0.669 | 0.612 | 0.085 |
| Port | 10 | 60 | 4.3 | 12 | 0.594 | 0.600 | −0.010 |
| Rata | 23 | 64 | 3.8 | 28 | 0.576 | 0.571 | 0.010 |
Micro-Checker detected excess of homozygotes for loci Rr14, Rr114 and D12 which can indicate presence of null alleles. Also, departure from Hardy-Weinberg equilibrium (HWE) was detected for those same loci (Table 2). Downstream analysis with and without these loci was performed, and since their presence did not show a significant effect, the following results are presented with all 14 loci included.
Number of alleles (Na), allelic richness (Ar; based on min. sample size of 10 individuals), observed heterozygosity (Ho) and within population gene diversity (Hs) according to Nei (1987), FST and FIS following Weir and Cockerham (1984) for each locus (*** indicates significance p < 0.001, based on 21,000 randomisations).
| Locus name | Na | Ar | Ho | Hs | Fst | Fis |
|---|---|---|---|---|---|---|
| Rr14 | 10 | 5.502 | 0.388 | 0.47 | 0.569*** | 0.231*** |
| D11 | 11 | 6.555 | 0.692 | 0.733 | 0.119*** | 0.068 |
| Rr114 | 9 | 5.231 | 0.437 | 0.574 | 0.035*** | 0.327*** |
| D7 | 9 | 6.683 | 0.699 | 0.665 | 0.251*** | −0.039 |
| D2 | 9 | 6.345 | 0.824 | 0.725 | 0.166*** | −0.129 |
| D18 | 13 | 8.34 | 0.725 | 0.767 | 0.171*** | 0.068 |
| D5 | 9 | 6.223 | 0.759 | 0.681 | 0.225*** | −0.14 |
| D20 | 14 | 6.464 | 0.706 | 0.763 | 0.029*** | 0.055 |
| D12 | 5 | 3.204 | 0.232 | 0.376 | 0.189*** | 0.411*** |
| D16 | 7 | 5.788 | 0.708 | 0.634 | 0.312*** | −0.109 |
| D15 | 10 | 6.771 | 0.591 | 0.669 | 0.261*** | 0.062 |
| D10 | 8 | 6.134 | 0.64 | 0.634 | 0.236*** | 0.024 |
| D19 | 10 | 6.564 | 0.593 | 0.593 | 0.346*** | 0.013 |
| Rr17 | 4 | 3.524 | 0.331 | 0.309 | 0.623*** | −0.064 |
| Average | 5.952 | 0.595 | 0.614 |
The pairwise Fst values suggest moderate genetic differentiation between the populations from the main island, Quixaba and Port (Fst = 0.1073, p < 0.01), and suggest very great differentiation between those two populations and Rata (Fst = 0.2956 and Fst = 0.3154, p < 0.01) (Table 3).
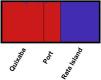
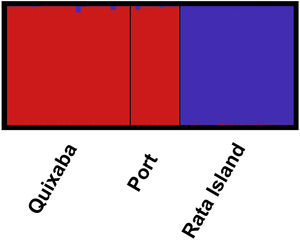
The great differentiation between Rata and the two populations from the main island (Quixaba and Port) was confirmed by the Bayesian clustering analysis performed in STRUCTURE (Fig. 3). This analysis, however, did not show differentiation between Quixaba and Port, indicating the most likely number of k = 2 (Supplementary Material 2).
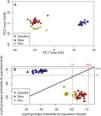
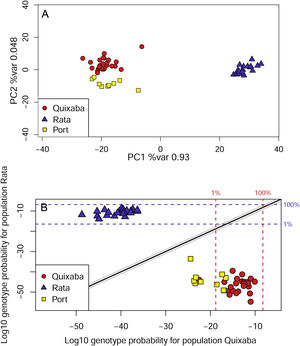
The genetic clustering in PCA (Fig. 4A) grouped Quixaba and Port individuals while Rata’s individuals formed an isolated group. The saddlepoint approximation method used in GenePlot (Fig. 4B) corroborated with PCA, STRUCTURE and Fst values indicating, a strong genetic differentiation between the populations of the two islands. The assignment of the Port population showed good fit to the Quixaba population for five individuals, meaning they are probably a subset of this population, while the other five individuals had poor fit to either population, and clustered with one individual from Quixaba that had poor fit to that population too. Overall, all analyses support genetic differentiation between the two island populations.
DiscussionAll results indicate strong population differentiation between the main island and Rata Island, with no evidence of recent gene flow between both, indicating that eradication on Rata Island has a high probability to be sustained over time.
Invasion historyThe presence of rats was officially reported on Fernando de Noronha by Branner in, 1888, but their introduction most likely occurred in the 16th century (Carleton and Olson, 1999) when European ships, mostly from Portugal and France, started to berth in the archipelago (Silva, 2013b). Over the years, the location was visited by ships from other European countries which could have led to multiple invasion events on the main island but this is not supported by D-Loop data. Mitochondrial sequence data revealed only two haplotypes each presented on a different island, probably because of some priority effect that happens when an established population prevents new immigrant colonisation (Fraser et al., 2015; Waters et al., 2013) or even due to genetic drift that might have caused loss of haplotypes and fixed the reported ones (Frankham et al., 2010). This pattern of a small number of introductions was also observed in larger scale in New Zealand, for R. rattus and R. norvegicus (Russell et al., 2019), and the Falkland Islands, for R. norvegicus (Hingston et al., 2016).
The difference between the two haplotypes was one base pair which suggests one could have derived from the other, but if this mutation originated in the Noronha archipelago we would expect to see one of the islands presenting both haplotypes; the original and the mutated one, which did not occur. The alternative hypothesis is that each island was colonised in independent events of invasion that brought with them different haplotypes. Both haplotypes are recorded in other islands around the world, and were the two most common registered in New Zealand (Russell et al., 2019) (GenBank MH751483 and MH751484). Hap1 was also reported by Colangelo et al. (2015) as the most common haplotype in the Mediterranean basin. Another sequence similar to Hap1 was reported by Hingston et al. (2005) as “HaMI” (Haplotypes absent from Madagascar and India) and belonged to New York, Great Britain, France and French Polynesia locations. Unfortunately, the provided sequence for HaMI ended at position 15,769 while our fragment ended at 15,925 so we could not compare 156 bp to be sure that these are the exactly same haplotypes. Independent colonisations seem most likely, however, we are aware that our limited number of samples and coverage may have camouflaged a rare individual with a different haplotype on either island.
Rat eradicationFernando de Noronha is located far from the coast and access to the archipelago is regulated, presenting a good potential for rat eradication. Still, even though most of the access to the archipelago happens by air on regular domestic aircrafts, there is heavy traffic of vessels and commodities between continental Brazil and the archipelago, promoting an infestation route for rodents. Fernando de Noronha's satellite islands, however, are inaccessible to the general public, and visitation by researchers is only allowed escorted by the National Park staff and with their vessel. Moreover, these secondary islands are important breeding sites for most marine seabirds (Mancini et al., 2016) and endemic species as they are free of other predators such as cats and possibly the tegu lizard (Abrahão et al., 2019). If rat eradication could be achieved on these satellite islands, re-invasion that is human mediated (i.e. vessel hitchhiking) would be unlikely, and a surveillance program could keep these rat-free. Moreover, the knowledge gathered during such programs could inform eradication actions for the main inhabited island, since this is not a trivial task. Failure rate of rat eradication on tropical islands is 2–2.5 times higher than on islands located in high latitudes (Russell and Holmes, 2015) and eradication on inhabited islands requires extra considerations (Oppel et al., 2011). In this study, we have investigated the feasibility of sustaining Rata Island rat-free after eradication, following the recommendation proposed by Russell et al. (2018). The results indicated no recent gene flow occurring between islands, which leads us to be optimistic about eradication sustainability on Rata Island,
In the Galapagos archipelago Willows-Munro et al. (2016) found limited admixture (significant pairwise Fst values) even between islands belonging to the same proximate region, but in that scenario the closest islands did not shown Fst values as high as we found between Rata Island and the main island. In fact, islands further apart from each other like San Cristobal and Isabela (130 km) were more closely related than Rata Island and the main island here are. On Fernando de Noronha, despite the fact that the north-east island chain suggests a stepping stone model of gene flow we found very great differentiation between Rata Island and the two populations from the main island (Quixaba and Port) (Fst > 0.25) indicating presence of very low or no current gene flow between the islands, which is corroborated by the partition of mitochondrial haplotypes, found to be exclusive to each island tested. This lack of gene flow, although surprising, can be explained by the topography of the islands Rasa, Sela Ginete, Meio and Rata. All four have cliffs on their borders (Silva, 2013a) which can be a significant factor in limiting gene flow for R. rattus, even with its climbing skills, as reported by Fewster et al. (2011). Besides there are three marine channels between the main island and Rata Island with strong water currents that could further make difficult the movement of rats between islands. Another possible explanation is that Sela Ginete is actually blocking gene flow between Rasa and Meio Islands because of its precipitous feature that can make it impracticable for rats to settle and disperse from there. In this case, it is possible that some gene flow occurs between Rasa Island and the main island and did occur prior to eradication between Meio Island and Rata Island. Unfortunately, we were not able to obtain samples from Sela Ginete due to its inaccessibility, if rats even exist there at all, and no samples were available from Meio Island prior to the rats being eradicated.
In contrast, the populations from Quixaba and Port, even being more distant from each other, present no barrier to dispersion besides distance and show only moderate differentiation according to Fst, suggesting presence of gene flow at some level. In the saddlepoint approximation method five Port individuals were shown as a subset of Quixaba and the other five individuals were assigned below the 1% line for the Quixaba population, meaning that even inside the same island there is some population structure. Besides, one individual from Quixaba was assigned below the 1% line for the Quixaba population as well, being grouped with the others from the Port and indicating probably an immigrant from the Port population.
RecommendationsComplete eradication in the archipelago is still not feasible due to legal restrictions and limited conservation funding (Russell et al., 2018). Nevertheless, our findings suggested a low probability of a reinvasion on Rata Island occurring from the main island, notwithstanding possible priority effects blocking current dispersal. We were able to define at least two eradication units for Fernando de Noronha, which following a successful rat eradication on Meio Island (unpublished data) encourages the execution of a Rata Island eradication as proposed by Russell et al. (2018). A rat eradication associated with biosecurity measures to prevent new invasions by hitch hiking on boats could create a sanctuary to protect and restore breeding seabird colonies that are known to be in decline (Mancini et al., 2016).
Data accessibilityAdditional information can be found in the electronic supplementary material. The microsatellite data file is available in the Figshare digital repository and can be accessed at: https://doi.org/10.17608/k6.auckland.11307497.v1.
PVE/CAPES 235453 & PDSE/CAPES 88881.189218/2018-01 from the Ministério da Educação, Brazil with Licence 43589-4 from ICMBio to research in Fernando de Noronha ICMBio/GEF-Mar and Vibha Thakur for the lab assistance in New Zealand. This study was financed in part by the Coordenação de Aperfeiçoamento de Pessoal de Nível Superior - Brasil (CAPES).